Viperin
Description
Viperins, for "Virus Inhibitory Protein, Endoplasmic Reticulum-associated, INterferon-inducible", are antiviral enzymes whose expression is stimulated by interferons in eukaryotic cells. They are important components of eukaryotic innate immunity and present antiviral activity against a wide diversity of viruses, including double-stranded DNA viruses, single-strand RNA viruses and retroviruses (N/A) .
Recently, Viperin-like enzymes were found in prokaryotes (pVips). Strikingly, like their eukaryotic counter-part with eukaryotic viruses, pVips provide clear protection against phage infection to their host and therefore constitute a new defense system (N/A) . Like eukaryotic Viperins, pVips produce modified nucleotides that block phage transcription, acting as chain terminators. They constitute a form of chemical defense. A recent study reported that pVips can be found in around 0.5% of prokaryotic genomes (N/A) .
Molecular mechanism
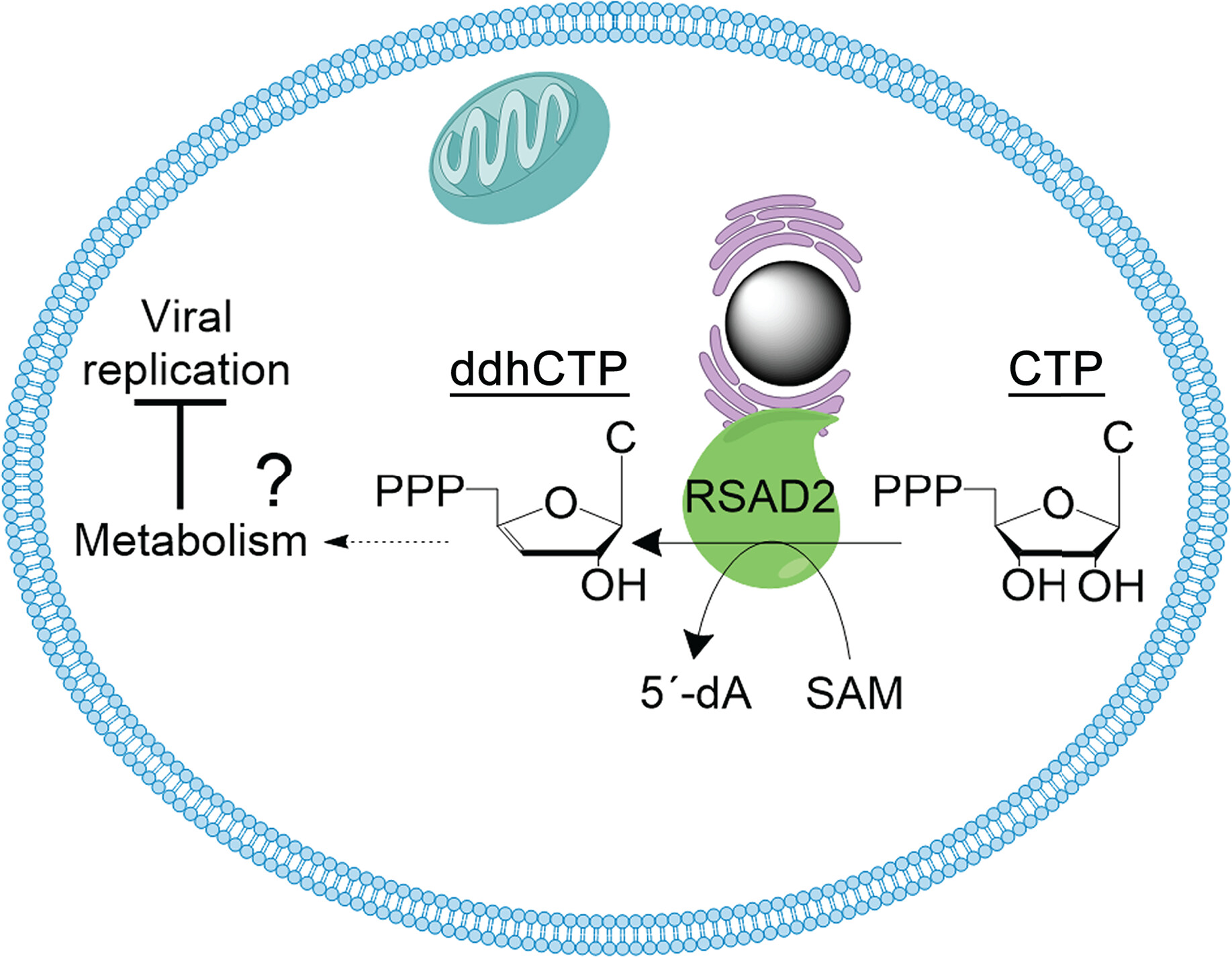 Fig.1: Catalytic activity of human Viperin generates ddhCTP (N/A)
Fig.1: Catalytic activity of human Viperin generates ddhCTP (N/A)
Viperins are members of the radical S-adenosylmethionine (rSAM) superfamily. This group of enzymes uses a 4Fe-4S] cluster to cleave S-adenosylmethionine (SAM) reductively, generating a radical that is generally transferred to a substrate. It was demonstrated that through their 4Fe-4S cluster catalytic activity, eukaryotic viperins convert a ribonucleotide, the cytidine triphosphate (CTP) into a modified ribonucleotide, the 3'-deoxy-3',4'-didehydro-CTP (ddhCTP) (N/A) .
Prokaryotic Viperins also convert ribonucleotides triphosphate into modified ribonucleotides, but contrary to their eukaryotic counterparts can use a diversity of substrates to produce ddhCTP, or ddh-guanosine triphosphate (ddhGTP), or ddh-uridine triphosphate (ddhUTP), or several of these nucleotides for certain pVips (N/A) .
Compared to the initial ribonucleotide triphosphate, the modified ddh-nucleotide product of Viperins lacks a hydroxyl group at the 3′ carbon of the ribose (Fig.1). The ddh-nucleotides produced by Viperins can be used as substrates by some viral RNA polymerases. Because of their lost hydroxyl group at the 3’carbon of the ribose, once incorporated into the newly forming viral RNA chain, these ddh-nucleotides act as chain terminators. By preventing further polymerization of the viral RNA chain, ddh-nucleotides can inhibit viral replication (N/A, N/A) .
Example of genomic structure
The Viperin is composed of 1 protein: pVip.
Here is an example found in the RefSeq database:
The Viperin system in Vibrio sp. SCSIO 43009 (GCF_023716525.1, NZ_CP071842) is composed of 1 protein: pVip (WP_182016311.1)